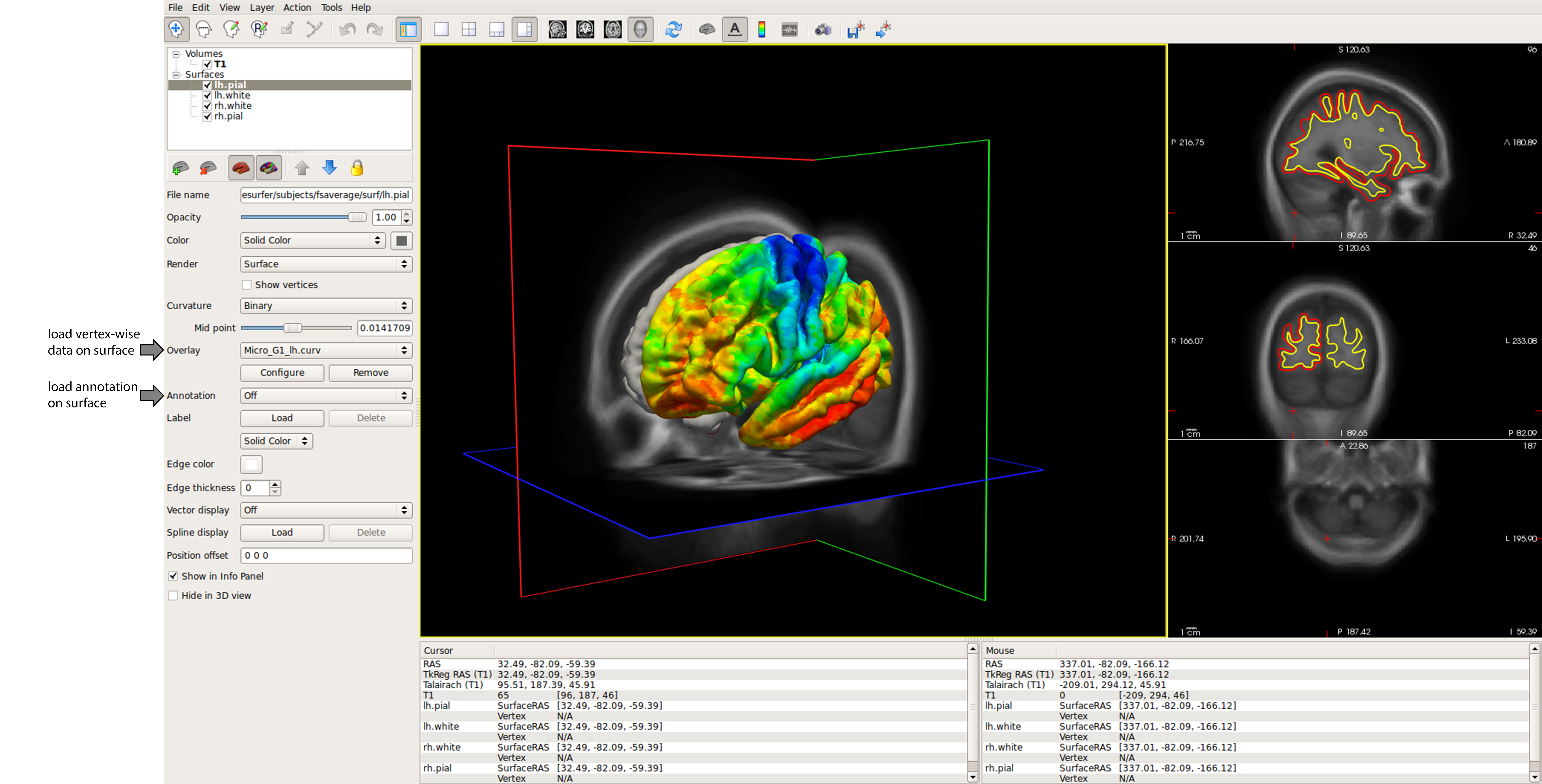
Visualisation across file formats¶
We should always visually check the fidelty of our data transformations. BigBrainWarp accepts and produces a range of file format, which require different types of visualisation. Here are a couple of tips for data visualisation, including a non-exhaustive list of visualisation tools for each file format.
| File format | Type | Visualisation tools |
|---|---|---|
| Nifti (.nii) | Volume | freeview, wb_view, MANGO, SurfStat |
| MINC (.mnc) | Volume | Display, MANGO, SurfStat |
| Gifti (.gii) | 3D geometry of surface (.surf.gii), vertex-wise data (.shape.gii) or vertex-wise labels (.label.gii) | wb_view, MANGO, Matlab |
| Wavefront obj (.obj) | 3D geometry of surface | SurfStat, Display |
| Freesurfer surface (.pial, .inflated, .white) | 3D geometry of surface | SurfStat, freeview |
| Curv (.curv) | Vertex-wise data | freeview, SurfStat |
| Annotation (.annot) | Vertex-wise labels and colour look up table | freeview, SurfStat |
Surfaces: Matlab-based, SurfStat-powered¶
Our favourite tool for surface-based analyses is SurfStat. SurfStat is a Matlab toolbox for statistical analysis and visualisation of neuroimaging data. You can download the toolbox as well as tutorials and extra helper functions from the micaopen github. By combining this with in-built Matlab functions, Freesurfer’s Matlab add-ons and the gifti toolbox, we can load and inspect MINC (.obj, .mnc), Freesurfer and Gifti datatypes in one easy space.
% set directories
GH = '/path/to/your/github/directories/
bbwDir = [GH '/BigBrainWarp/'];
% add directories to path
addpath(genpath([GH '/micaopen/surfstat']));
addpath([bbwDir '/scripts/']);
addpath(genpath('/path/to/gifti-1.6/')); % download from https://www.artefact.tk/software/matlab/gifti/
% 1. Load the surface
% .obj
BB = SurfStatAvSurf({[bbwDir '/tpl-bigbrain/tpl-bigbrain_hemi-L_desc-pial.obj'], ...
[bbwDir '/tpl-bigbrain/tpl-bigbrain_hemi-R_desc-pial.obj']});
% Binary Freesurfer surfaces (ie: .pial, .white, .sphere etc)
FS = SurfStatAvSurf({[bbwDir '/tpl-fsaverage/tpl-fsaverage_hemi-L_desc.pial'], ...
[bbwDir '/tpl-fsaverage/tpl-fsaverage_hemi-R_desc.pial']});
% Gifti surfaces
tmp_lh = gifti([bbwDir '/spaces/tpl-fs_LR/tpl-fs_LR_hemi-L_den-32k_desc-inflated.surf.gii']);
tmp_rh = gifti([bbwDir '/spaces/tpl-fs_LR/tpl-fs_LR_hemi-R_den-32k_desc-inflated.surf.gii']);
FSLR32.coord = [tmp_lh.vertices' tmp_rh.vertices']; % coord and tri are the two expected components of a SurfStat surface structure
FSLR32.tri = [tmp_lh.faces; tmp_rh.faces+length(tmp_lh.vertices)];
% 2. Load the data
% .txt data
tmp_lh = readmatrix([bbwDir '/spaces/tpl-bigbrain/tpl-bigbrain_hemi-L_desc-Hist_G1.txt']);
tmp_rh = readmatrix([bbwDir '/spaces/tpl-bigbrain/tpl-bigbrain_hemi-R_desc-Hist_G1.txt']);
BB_G1 = [tmp_lh; tmp_rh];
% Freesurfer-style curv data
tmp_lh = read_curv([bbwDir '/spaces/tpl-fsaverage/tpl-fsaverage_hemi-L_den-164k_desc-Micro_G1.curv']);
tmp_rh = read_curv([bbwDir '/spaces/tpl-fsaverage/tpl-fsaverage_hemi-R_den-164k_desc-Micro_G1.curv']);
FS_G1 = [tmp_lh; tmp_rh];
% gifti data
tmp_lh = gifti([bbwDir '/spaces/tpl-fs_LR/tpl-fs_LR_hemi-L_den-32k_desc-Hist_G1.shape.gii']);
tmp_rh = gifti([bbwDir '/spaces/tpl-fs_LR/tpl-fs_LR_hemi-R_den-32k_desc-Hist_G1.shape.gii']);
FSLR32_G1 = [tmp_lh.cdata; tmp_rh.cdata];
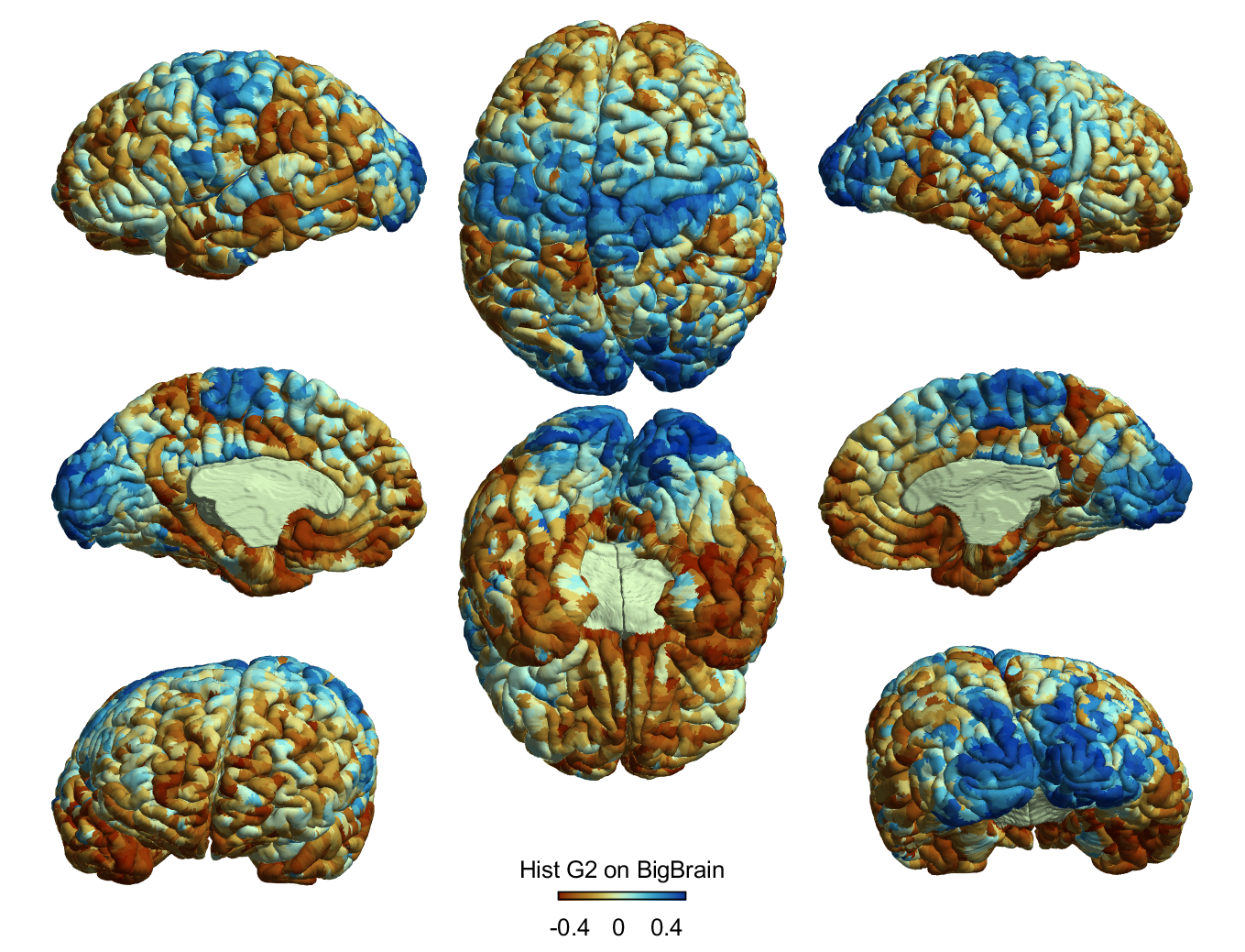
% 3. Visualise data
figure; SurfStatViewData(BB, BB_G1)
figure; SurfStatViewData(FSLR32, FSLR32_G1)
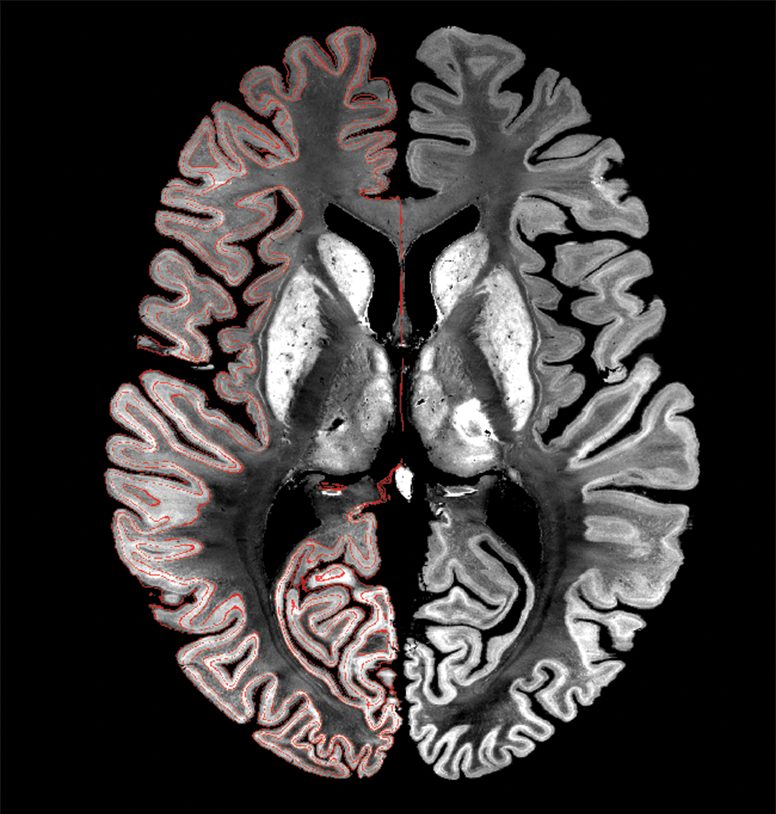
Displaying MINC¶
The volume-based transformations in BigBrainWarp depend upon MINC. BigBrainWarp enables conversions to nifti, so it may not be necessary to check the intermediary .mnc files yourself, but if you would like to then we’ll turn your attention towards Display. With Display, you can also overlay .obj surfaces on the volume.
Display volume_file.mnc
# Click "File" then "Load File"
# in the terminal
/full/path/to/surface_file.obj
# Return to Main Menu, Click "Objects" then "Write Object to File"
MANGO 🥭¶
MANGO is a Multi-Image Analysis GUI that supports a wide range of imaging file formats. Bonus, it is very easy to install and run on any operating system (http://mangoviewer.com/index.html).
Freeview¶
Freeview is the built-in visualisation tool of Freesurfer and is handy for all Freesurfer-style file formats.